Bases2FASTQ: Setting up for FASTQ generation with AWS
- AUGenomics .
- Jun 6, 2024
- 3 min read
Welcome to our Element AVITI Users' Hub by AUGenomics! As early AVITI adopters, we understand that sometimes finding simple answers using new technology isn't so simple. That's why we've decided to share knowledge in a way that we wish existed when we adopt new tech. Enjoy!

So, you just got your first AVITI.. or maybe you've received a raw AVITI run folder to analyze. Now what? When sequencing with the Element AVITI, you'll quickly notice that FASTQ generation doesn't use the typical bcl2fastq program you may be used to by Illumina. Instead, you'll need to use bases2fastq released by Element Biosciences.
If you're used to a user-friendly, no-code platform like Basespace, your first time using bases2fastq may feel like a bit of a learning curve since this FASTQ generation is done through a cli rather than a cloud-based GUI/web app. This step-by-step guide will help you with the basics of setting up and running bases2fastq through AWS EC2 using AWS S3 storage so that you can get your demultiplexing done in a pinch.
In this guide, we assume raw run folders are stored on an AWS S3 bucket. From there, an AWS EC2 can pull data from the S3 bucket, process the run through a temporary folder, and transfer data to the S3 bucket of your choice.
To start, click "Launch Instance" from the AWS EC2 application:

Use the following Image Settings:
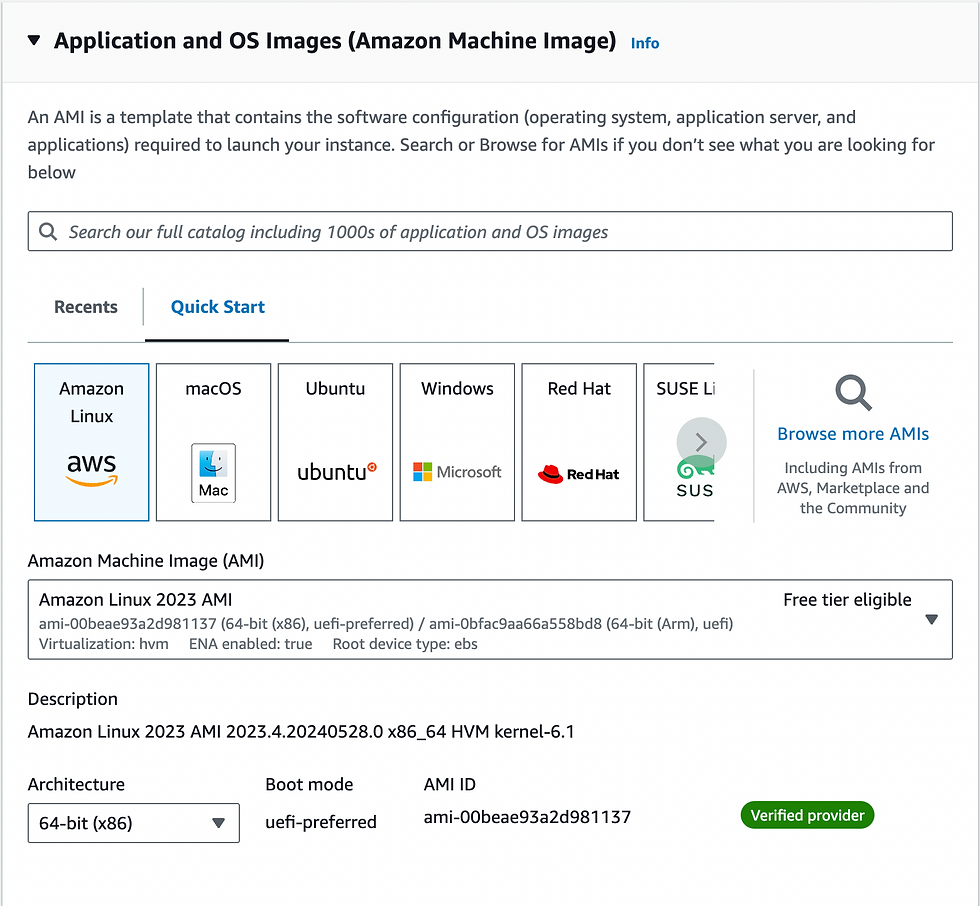
Choose your instance type:
At AUGenomics, we use an m5dn.12xlarge EC2 Instance with 800GiB for bases2fastq processing. For optimal performance, make sure you have at least 16 CPU cores available and enable threading Bases2Fastq with the -p argument. The following memory requirements apply to both Docker and static binary distributions:
A 2 x 75 or 2 x 150 AVITI System run requires 4 GB RAM per concurrent thread.
A 2 x 300 AVITI System run requires 6 GB RAM per concurrent thread.

Set your storage:
When using cloud storage, Bases2Fastq downloads input files and stages output files in a temporary directory. Intermediate files generated during analysis are also stored in the temporary directory. After an execution completes, the temporary directory is cleared.
The temporary directory typically uses 400–500 GB for a 2 x 150 run. For some applications, a run can use up to 800 GB. The necessary amount of scratch space depends on the number of polonies and cycles in the run and the optional arguments in the Bases2Fastq execution.

2. Launch and connect to your instance
This can be done quickly through EC2 Instance Connect on your browser. For long term usage, we recommend connecting through SSH using your computer's terminal. Once connected, you'll be executing the remaining steps through the EC2's shell.
3. Install bases2fastq and all necessary dependencies on your fresh EC2
sudo yum update
sudo yum install docker
sudo yum install sysstat parallel pigz -y
sudo yum install tree -y
4. Configure your credentials and setup docker service export AWS_ACCESS_KEY_ID=[Your_Access_Key]
export AWS_SECRET_ACCESS_KEY=[Your_Secret_Access_Key]
export AWS_DEFAULT_REGION=[Your_Region]
eval "$(aws configure export-credentials --profile default --format env)"
sudo service docker start
sudo usermod -a -G docker ec2-user
mkdir output
sudo mount -o remount,size=800G,noexec,nosuid,nodev,noatime /tmp
4. Run bases2fastq
sudo docker run -it -v "${PWD}/output":"/output/" -e AWS_ACCESS_KEY_ID=$AWS_ACCESS_KEY_ID -e AWS_SECRET_ACCESS_KEY=$AWS_SECRET_ACCESS_KEY -e AWS_DEFAULT_REGION=$AWS_DEFAULT_REGION elembio/bases2fastq:latest bases2fastq -p 45 s3://[RunFolderLocation] /output/ -r s3://[RunManifestLocation].csv
6. Congratulations! You can now transfer your FASTQ files and demux logs back to your S3
cd output
aws s3 sync . s3://[YourBucketDirectory]
Element Biosciences has recently released automated workflows through their cloud-based platform. These have some advantages and drawbacks depending on your specific sequencing pipeline and usage. Check back in for reviews and user notes as we'll be posting more tips and tricks for using the AVITI platform!
Happy Analyzing,
AUG Team

Comments